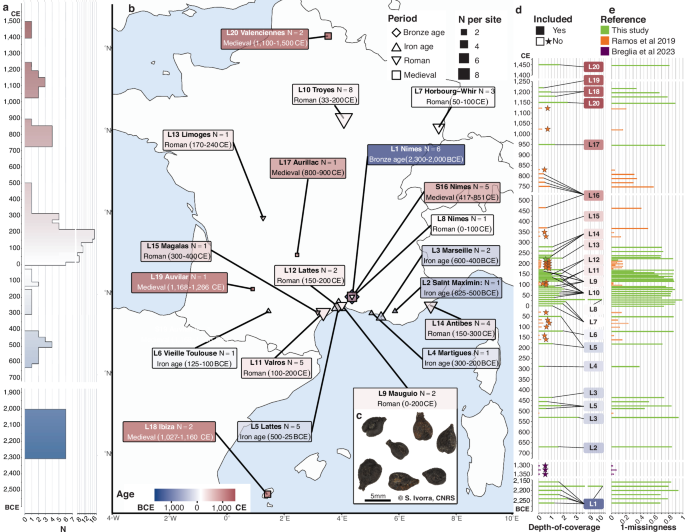
Genome-wide ancient DNA from 49 archaeological grape pips spanning ~4000 years shows domesticated grape use in France by ~625–500 BCE and widespread genetic diversity introduced during the Roman period. Vegetative (clonal) propagation is evident by the mid-Iron Age, with one Medieval Valenciennes pip genetically identical to modern 'Pinot Noir', demonstrating clonal continuity of roughly 600 years. Results highlight deep genetic continuity and long-distance exchange shaping modern French viticulture, with limited direct near-term market impact beyond cultural/heritage and branding considerations for wine producers.
This dataset makes clear a structural demand shift: projects are moving from low-marker panels to whole-genome shotgun workflows that require far higher sequencing throughput, more upstream sample prep, and routine QC instrumentation. That favors suppliers with dominant flowcell economies and integrated consumable franchises — every additional gigabase per sample translates into recurring revenue for flowcells, extraction kits, and QC assays over multi-year archaeological programs. Second-order winners will be companies embedded in the front-end wet lab and QC stack (sample extraction, library prep, fragment analysis) because repeated re-sequencing, imputation-driven rescues of low-coverage data, and multi-site collaborations lock in consumable cadence; conversely, vendors whose revenue depends on targeted assay sales only face substitution risk. Geopolitical and ethical constraints around cross-border sample movement create demand for regional sequencing hubs and on-site instrumentation, shifting procurement toward vendors with strong EU supply/resale footprints and field-deployable QC tools. Near-term catalysts: funding cycles and large consortia grants that underwrite multi-year sequencing commitments, plus methodological trends (imputation and ancient haplotype panels) that convert low-coverage datasets into publishable assets — both boost spend per sample months-to-years out. Tail risks include a rapid, cost-disruptive breakthrough in a competing long-read or nanopore modality that undercuts incumbent flowcell economics, and regulatory limits on destructive sampling or export of cultural heritage specimens that would shrink addressable projects in weeks-to-months. My base-case: secular growth in high-value niche genomics (archaeogenomics, heritage/agri‑omics) will be a modest but durable add-on to human/clinical sequencing demand over 12–36 months, concentrated in consumables and mid-range instrumentation rather than one-off capital explosions; positioning should focus on recurring consumable capture and regional deployment resilience.
AI-powered research, real-time alerts, and portfolio analytics for institutional investors.
Request a DemoOverall Sentiment
neutral
Sentiment Score
0.00
Ticker Sentiment